All published articles of this journal are available on ScienceDirect.
Prevalence of β-lactam (blaTEM) and Metronidazole (nim) Resistance Genes in the Oral Cavity of Greek Subjects
Abstract
Objectives:
The aim of this study is to investigate the prevalence of blaTEM and nim genes that encode resistance to β-lactams and nitroimidazoles, respectively, in the oral cavity of systemically healthy Greek subjects.
Materials and Methodology:
After screening 720 potentially eligible subjects, 154 subjects were recruited for the study, including 50 periodontally healthy patients, 52 cases of gingivitis and 52 cases of chronic periodontitis. The clinical parameters were assessed with an automated probe. Various samples were collected from the tongue, first molars and pockets >6mm, and analysed by polymerase chain reaction-amplification of the blaTEM and nim genes, using primers and conditions previously described in the literature.
Results:
There was a high rate of detection of blaTEM in plaque and tongue samples alike in all periodontal conditions (37% of plaque and 60% of tongue samples, and 71% of participants). The blaTEM gene was detected more frequently in the tongue samples of the periodontally healthy (56%) and chronic periodontitis (62%) groups compared to the plaque samples from the same groups (36% and 29%, respectively; z-test with Bonferroni corrections-tests, P<0.05). The nim gene was not detected in any of the 343 samples analysed.
Conclusion:
The oral cavity of Greek subjects often harbours blaTEM but not nim genes, and therefore the antimicrobial activity of β-lactams might be compromised.
INTRODUCTION
Antimicrobial resistance has emerged as a major global issue, compromising the effective treatment of potentially life-threatening infections [1]. Resistance against important medical pathogens has been recognised and investigated worldwide, however, there is limited data in the literature concerning resistance in the oral cavity [2].
Various classes of antimicrobials, including several administered for periodontal infections, are ineffective against bacterial species due to the acquisition of antibiotic resistance, which is frequently spread by genetic material [3]. Antimicrobial resistance genes in the oral environment can be transferred among different species, or even genera, of bacteria by conjugation or plasmids, and can regulate various mechanisms of resistance [4, 5].
Typical examples of β-lactam resistance mechanisms are the production of antibiotic-inactivating enzymes, such as the β-lactamases, and alterations in the β-lactam targets, that is the penicillin-binding proteins (PBPs) [6]. Another antibiotic resistance mechanism is the enhanced function of efflux pumps, which are known to transport antimicrobial molecules outside the bacterial cell [7]. In the case of nitroimidazoles, apart from the enhanced function of efflux pumps, another significant mechanism is the deficient prodrug activation, through alternative metabolic pathways, resulting in non-toxic compounds [8].
It has been shown that resistance to antibiotics is directly correlated to their usage for both antimicrobial treatment and other purposes (such as growth promotion in animals for aquaculture and agriculture), and, therefore, to the related policy of each country [9]. β-Lactams and metronidazole are commonly prescribed for dental infections and, thus, the prevalence of resistance to these antibiotics may have clinical implications. The purpose of the present study was to investigate the prevalence of blaTEM and nim genes, which contribute to β-lactam and metronidazole resistance, respectively, in the oral cavity of Greek subjects with various periodontal conditions.
MATERIALS AND METHODOLOGY
Subject Sample and Study Design
From September 2011 to April 2014, 720 subjects attending the Clinic of Periodontology at 251 Air Force Hospital, Athens, Greece and the Clinic of the Department of Preventive Dentistry, Periodontology and Implant Biology, Dental School, Aristotle University of Thessaloniki, Greece, were screened for enrolment in the study.
Inclusion Criteria
To be included in the study, subjects should fulfill the following criteria:
- a) Age > 30 years.
- b) Absence of systemic diseases affecting the subjects’ immune system or medications known to affect perio-dontal tissues (causing gingival enlargement), infectious conditions (hepatitis, HIV) or pregnancy and lactation.
- c) No periodontal treatment or antibiotic intake during the last six months.
In addition, patients were required to meet one of the following conditions:
- a) Periodontal health (bleeding on probing <10%, without any clinical attachment loss ≥3 mm).
- b) Gingivitis (bleeding on probing >20%, without any clinical attachment loss ≥3 mm), or
- c) Moderate to advanced chronic periodontitis (presence of proximal attachment loss of ≥5 mm in ≥30% of teeth present) [10].
A total of 154 patients were finally enrolled in this cross-sectional study which were divided into three groups as follows: a) periodontally healthy (n = 50), b) gingivitis (n = 52), and c) chronic periodontitis (n = 52).
The study was conducted according to the protocol outlined by the Research Committee, Aristotle University of Thessaloniki, Greece and approved by the Ethical Committee of the School of Dentistry (#120), while it was also in compliance with the ethical principles of the World Medical Association Declaration of Helsinki. All patients read and signed an approved informed consent document prior to participation in the study.
Clinical Assessments
Clinical recordings were performed at six points (mesio-buccal, buccal, disto-buccal, mesio-lingual, lingual, and disto-lingual). Those were performed by a calibrated examiner (GK) routinely performing periodontal examination and therapy at 251 General Airforce Hospital using an automated probe (Florida probe, Florida Probe Corporation, Gainesville, FL, USA). The automated probe is documented for accuracy and reliability [11].
The recordings included the following parameters:
- a) Probing Pocket Depth
- b) Recession
- c) Clinical Attachment Level
- d) Bleeding on Probing
Clinical Sampling
Two clinical samples were collected from each patient: a pooled subgingival plaque sample from the mesio-buccal surface of the four first molars (or premolars in the absence of molars), taken with sterile Gracey curettes; and a sample collected from the dorsal surface of the tongue with a sterile straight surgical bone curette (Sklar Instruments, PA, USA), after applying three consecutive strokes. All samples were immediately placed in 200 μl of TE buffer (Tris HCL 10 mM, EDTA 1 mM, pH 7.5) and stored at -20°C until assayed. In addition, a third clinical sample, taken from the deepest periodontal pocket, was collected from chronic periodontitis patients.
Polymerase Chain Reaction (PCR)
The Analysis of these specimens was performed blindly on coded samples. All experiments were carried out in the Department of Microbiology, School of Biology, Aristotle University of Thessaloniki, Greece. DNA isolation was achieved by using a commercial DNA isolation kit (Qiagen, QIAamp DNA Mini Kit, Qiagen, Hilden, Germany). PCR was initially used for the detection of the 16S ribosomal RNA gene to verify that the clinical samples contained identifiable bacterial DNA [12]. The samples were further analysed by PCR for the presence of genes that confer resistance to β-lactams and metronidazole. The blaTEM gene was chosen as an initial screening test, because TEM is the leading gene conferring resistance to β-lactams and, through mutations, it leads to the emergence of extended spectrum β-lactamases (ESBLs) [13]. PCR conditions and primers are described in Table 1 [14, 15]. A Peltier Thermal Cycler (PTC-100, Peltier Thermal Cycler, MJ Research) was used for PCR.
Polymerase chain reaction (PCR) conditions for 16S, blaTEM and nim.
| PCR conditions | 16S (625 bp) | blaTEM (618 bp) | nim (458 bp) | |
| Initial denaturation | 95°C for 5 min | 94°C for 5 min | 94°C for 10 min | |
| Denaturation | 94°C for 1 min | 94°C for 1 min | 94°C for 30 s | |
| Annealing | 55°C for 1 min | 62°C for 1 min | 62°C for 1 min | |
| Extension | 72°C for 1 min | 72°C for 1 min | 72°C for 1 min | |
| Step 2 to 4 | 34 cycles | 35 cycles | 32 cycles | |
| Final extension | 72°C for 5 min | 72°C for 10 min | 72°C for 10 min | |
| DNA template | 5 µl | 5 µl | 5 µl | |
| dNTPs mix, 10 mM | 1 µl (200 µM) | 2 µl (400 µM) | 1 µl (200 µM) | |
| Reaction Buffer 5x | 10 µl | 10 µl | 10 µl | |
| MgCl2 Solution, 25 mM | 3 µl (1.5 mM) | 4 µl (2 mM) | 3 µl (1.5 mM) | |
| Upstream Primer, 10 µM | 2.5 µl (0.5 µM) | 5 µl (1 µM) | 5 µl (1 µM) | |
| Downstream Primer, 10 µM | 2.5 µl (0.5 µM) | 5 µl (1 µM) | 5 µl (1 µM) | |
| Taq polymerase, 5 U/µl | 0.25 µl (1.25 U) | 0.25 µl (1.25 U) | 0.25 µl (1.25 U) | |
| Water for injection | 25.75 µl | 18.75 µl | 20.75 µl | |
| Values in parenthesis are final concentration | ||||
16S primers 5'-CAG GAT TAG ATA CCC TGG TAG TCC ACG C-3' and 5'- GAC GGG CGG TGT GTA CAA GGC CCG GGA ACG-3' [12] blaTEM primers 5'-AGA TCA GTT GGG TGC ACG AG -3' and 5'-CAG TGC TGC AAT GAT ACC GC -3' [15] nim primers 5'-ATG TTC AGA GAA ATG CGG CGT AAG CG-3' and 5'-GCT TCC TTG CCT GTC ATG TGC TC-3' [14]
Both negative and positive controls were used for each set of samples analysed by PCR. Sterile water for injection (Demo S.A. Pharmaceutical Industry) was used as a negative control (replacing DNA template in the PCR reaction mixture) and DNA from the BL21 strain of E. coli, infected with the PGRF plasmid known to carry the blaTEM gene (pGFPuv, Clontech labs, USA), and B. fragilis, carrying the nim gene, were used as positive controls.
The PCR amplification products were electrophoresed through a 2% agarose gel, stained with ethidium bromide, exposed under UV light and photographed. A 100 base pair (bp) DNA ladder (New England Biolabs) was used as a molecular weight standard. The amplified fragment sizes were 625 bp for 16S rRNA, 618 bp for blaTEM and 458 bp for nim. Gel electrophoresis was carried out twice for each PCR product to test the reproducibility of the results (Supplementary file 1). For verification purposes, random PCR products were sent for sequencing (VBC-Biotech, Vienna, Austria) and confirmed the identity of the amplicons.
Questionnaire
All subjects were interviewed by one of the authors (DS) and completed a questionnaire about the following:
a) Smoking.
b) Frequency of antibiotic intake for medical and dental reasons, class of antibiotics used 6-12 months before the
interview as well as up to 5 years prior to it.
c) Use of antibiotics without a prescription.
d) Availability of antibiotics at home.
e) Awareness of the problem of antibiotic resistance.
To avoid errors on behalf of the participants concerning the use of antibiotics, the class of antibiotics consumed was recorded based on the subjects’ personal national health record.
Statistical Analyses
The statistical analysis was carried out with the statistical package SPSS Statistics v.19 software (IBM, Armonk, NY, US). The experiment was set to have at least 90% power to detect changes of 25% with a significance level (alpha) of 0.05 (two-tailed). A pilot study provided the expected values of the primary outcome variable [16]. For clinical parameters, indicators of descriptive statistics were used, such as the mean and standard deviation for each group, with the patient as the observational unit. Discrepancies in clinical parameters were detected by applying the Kruskal-Wallis test. Variations based on the distribution of participants according to gender and smoking, were detected by applying the z-test for proportions, adjusted with Bonferroni corrections. Differences in the prevalence of the investigated genes and answers to the questionnaire among periodontally healthy, gingivitis and periodontitis subjects, and within each group, were detected by applying the z-test for proportions, adjusted with Bonferroni corrections.
RESULTS
Clinical parameters and demographic data for participants are summarized in (Table 2). All groups varied substantially in terms of clinical parameters of periodontal disease (Kruskal-Wallis test, P<0.05). Although participants did not present statistically significant differences regarding age and smoking status, male subjects outnumbered the female ones in the chronic periodontitis group (z-test with Bonferroni corrections, P<0.05).
Demographic data and clinical parameters.
| Diagnosis | Total | Age (mean years ± SD) | Male (%) | Smokers (%) | Probing depth (mm) (mean ± SD) | Recession (mean ± SD) | Clinical attachment level (mm) (mean ± SD) | Bleeding on probing (%) |
|---|---|---|---|---|---|---|---|---|
| Periodontal health |
50 | 46 ± 8 | 52 (a) |
24 | 1.54 ± 0.2 (a) |
0.04 ± 0.06 (a) | 1.38 ± 0.24 (a) | 6% ± 5% (a, b) |
| Gingivitis | 52 | 49 ± 9 | 42 (b) |
35 | 1.89 ± 0.3 (b) |
0.06 ± 0.16 (b) | 1.61 ± 0.4 (b) |
34% ± 15% (a) |
| Chronic periodontitis |
52 | 50 ± 10 | 79 (a, b) |
38 | 3.05 ± 0.6 (a, b) |
0.31 ± 0.42 (a, b) | 2.81 ± 0.6 (a, b) |
32% ± 21% (b) |
Statistically significant differences between groups are indicated by the same letter (Kruskal Wallis test and z-test with Bonferroni corrections, P < 0.05)
The distribution of blaTEM and nim is summarized in (Fig. 1). The nim gene was not detected in any of the 343 samples analysed. The overall presence of blaTEM in the subjects was estimated to be 71% (109 out of 154 subjects were positive in at least one sample).
The prevalence of blaTEM was similar in plaque or tongue samples for all three periodontal conditions (periodontal health, gingivitis and chronic periodontitis), when comparing across the same sampling site (z-test with Bonferroni corrections, P>0.05).
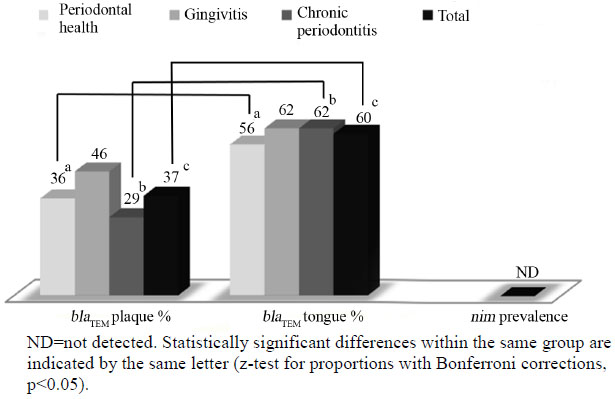
Prevalence of blaTEM (and nim) in plaque and tongue samples according to periodontal conditions.
The blaTEM gene was detected at a significantly higher rate in the tongue sample in both the periodontally healthy (56%) and the chronic periodontitis (62%) groups, compared to the plaque samples in these same groups (36% and 29%, respectively; z-test with Bonferroni corrections, P<0.05). In the chronic periodontitis subjects, the blaTEM gene was detected at a significantly lower rate in the deep periodontal pocket samples (26%) compared to the tongue samples (62%; z-test with Bonferroni corrections, P<0.05).
Gingivitis subjects, irrespective of the sampling site, had a significantly higher frequency of detection of the blaTEM gene (54%) compared to chronic periodontitis groups (40%)(z-test with Bonferroni corrections, P<0.05).
Table 3 summarizes the results from the questionnaire. The following major differences were observed among the groups: compared to periodontally healthy individuals, chronic periodontitis patients are less aware of antibiotic resistance and have been administered tetracyclines less frequently (P<0.05). β-Lactam antibiotics were the only class of antibiotics that had been administered during the 6-12 months before the survey and the most frequently prescribed antibiotics over the past 5 years (P<0.05) in all three groups of participants.
DISCUSSION
The current study was conducted in an attempt to investigate the frequency of detection of bacterial genes contributing to resistance to β-lactams and metronidazole in three locations in the oral cavity: subgingival plaque and tongue, from all patients, and deep periodontal pockets >6 mm from chronic periodontitis patients. Our results establish that one of the important genes encoding for the family of β-lactamases (blaTEM) is frequently detected in the oral cavity (tongue and plaque samples), regardless of the periodontal conditions (periodontal health, gingivitis or chronic periodontitis). Different periodontal conditions were assessed due to the fact that they reflect the presence of a wide variety of bacteria (predominance of anaerobes in periodontitis), which could lead to a difference in the presence of resistance genes. That being said, previous studies have assessed prevalence only in periodontitis patients [17, 18]. In our study gingivitis subjects had higher prevalence of resistance genes (54%) in comparison to periodontitis subjects (40%), suggesting that Gram-negative anaerobes might not be the main carriers of these genes. The sampling site was found to be more important than the periodontal condition, with the dorsal surface of the tongue consistently carrying the majority of the resistance genes regardless of the periodontal condition. In contrast, the gene conferring resistance to metronidazole (nim) was not detected in any of the 343 samples analysed in this study.
Consumption, attitude and knowledge about antibiotics of the subject sample.
| Periodontal health (%) N=50 |
Gingivitis (%) N=52 |
Chronic periodontitis (%) N=52 |
TOTAL (%) N=154 |
||
|---|---|---|---|---|---|
| Antibiotic use for medical reasons during the last 5 years (%) |
1-2 times | 38%(19/50) | 44%(23/52) | 46%(24/52) | 43% |
| 3-4 times | 26%(13/50) | 25%(13/52) | 29%(15/52) | 27% | |
| >5 times | 36%(18/50) (a) |
27%(14/52) | 15%(8/52) (a) |
26% | |
| Antibiotic use for dental reasons during the last 5 years (%) |
1-2 times | 54%(27/50) | 42%(22/52) | 42%(22/52) | 46% |
| 3-4 times | 8%(4/50) | 10%(5/52) | 8%(4/52) | 8% | |
| >5 times | 0% | 4%(2/52) | 6%(3/52) | 3% | |
| β-Lactams (%) | Yes | 89%(39/44) | 73%(37/51) | 75%(38/51) | 78% |
| tetracyclines (%) imidazoles (%) |
Yes | 31%(13/42) (a) |
20%(10/51) | 6%(3/47) (a) |
18% |
| Yes | 17%(7/41) | 24%(12/51) | 13%(6/47) | 18% | |
| Available at home (%) | Yes | 30%(15/50) | 33%(17/52) | 17%(9/52) | 27% |
| Without Prescription (%) | Yes | 2%(1/50) | 2%(1/52) | 0%(0/52) | 1% |
| By a pharmacist | 14%(7/50) | 25%(13/52) | 15%(9/52) | 18% | |
| Antibiotic intake last 6-12 months (%) |
Yes (β-lactams) | 28%(14/50) | 14%(7/51) (a) |
33%(17/52) (a) |
25% |
| Knowledge about antimicrobial resistance (%) |
Yes | 48%(24/50) (a) |
29%(15/51) | 23%(12/52) (a) |
33% |
Statistically significant differences among groups are indicated by the same letter. Statistically significant differences within each group are indicated by bold lettering (z-test for proportions with Bonferroni corrections, P<0.05).
β-Lactams are one of the most frequently prescribed classes of antibiotics for both medical and dental purposes, however, there is substantial concern about the compromised efficacy of these antimicrobials due to the development of bacterial resistance. Studies in the dental literature have addressed the issue of periodontal pathogen susceptibility to β-lactam antibiotics (mainly amoxicillin) and whether certain bacterial species, for example, members of the oral microbiota such as Prevotella spp., can contribute to the establishment and dissemination of bacterial resistance in the subgingival environment [17].
Studies in Europe, using cultures of bacteria grown on selective plates, showed an important difference between the Netherlands and Spain in the amoxicillin resistance of periodontal strains of bacteria, reflecting the antibiotic prescription policy of each country [19]. A recent large-scale study in the US investigated antimicrobial resistance in 400 chronic periodontitis patients and reported a frequent occurrence of resistant species of subgingival periodontal pathogens [18, 20]. In this study, 74.2% of the patients harboured species resistant to several antibiotics including doxycycline, amoxicillin, clindamycin and metronidazole, and 43.3% of the patients had one or more test species that were resistant to amoxicillin. In the same study, 30.3% of patients harboured one or more metronidazole–resistant species, which included, as expected, exclusively non-anaerobic test species, such as Streptococci spp. In this study only 4 samples from anaerobic bacteria were truly metronidazole resistant, as metronidazole prodrug can only be activated in anaerobic conditions and therefore aerobic bacteria are intrinsically resistant. These studies collectively suggest that subgingival pathogens currently include species that are resistant to β-lactam antibiotics and nitroimidazoles.
Several studies have focused on the identification of the bacterial genera and species in the oral cavity that are sources of β-lactamases, one of the major mechanisms conferring resistance to β-lactams [21-23]. According to a well-established report, several members of the genus Prevotella, and Fusobacterium nucleatum are considered major producers of these enzymes, in addition to other species such as Tannerella forsythia, Neisseria and Capnocytophaga spp., which have also been shown to produce these enzymes [24].
The outcome of the present study suggests, a correlation between the observed frequency of the blaTEM gene and the high intake of the β-lactam class of antibiotics. A temporary increase in mean amoxicillin-resistant isolates has also been observed in chronic periodontitis patients after mechanical treatment and adjunctive therapy with amoxicillin for 14 days, but the percentage of resistant strains returned to pre-treatment levels after 90 days [25]. A recent report has also shown that subjects with generalized aggressive periodontitis (GAP) treated with adjunctive metronidazole plus amoxicillin have shown a significant increase in anaerobic microbiota resistant to amoxicillin within 2 months, returning to baseline values after 6 months, and indicated possible risks for increasing antimicrobial resistance to the population when empirically prescribing amoxicillin [26]. Thus, in order to avoid overestimation of prevalence, according to the present criteria for inclusion, none of the participants has used systemic antibiotics at least 6 months prior to enrollment in the study.
The personal official national health record of participants during the last five years has shown that >90% have received a course of antibiotics for medical reasons and >56% for dental reasons, with β-lactams being the significantly predominant class prescribed (>78%). According to the 2013 European Centre for Disease Prevention and Control report, Greece had one of the highest rates of antibiotic consumption among the European countries (39.4 defined daily doses or DDDs per 1000 inhabitants), a parameter known to significantly affect antimicrobial resistance, and β-lactams were among the most frequently prescribed antibiotics (21.8 DDDs of which 12.9 were penicillin) [27].
In a previous study, regarding Greek subjects, the high prevalence of tetracycline resistance genes tetM (>92% of subjects) and tetQ (>75% of subjects) in the oral cavity could not be correlated to the intake of tetracyclines (18% of participants during the last 5 years) suggesting other sources of these specific genes, such as the environment or food chain [28]. In contrast, systemic administration of β-lactams appears to be a possible factor affecting the dissemination of the blaTEM gene in the present study population.
Metronidazole is one of the most popular antimicrobials in periodontology, due to its narrow spectrum against anaerobic bacteria, and has regained medical interest because of its effectiveness against Helicobacter pylori, the main cause of gastroduodenal ulcer, and against post surgical anaerobic infections. Bacterial resistance to metronidazole is usually conferred by the presence of nim genes, which encode a nitroimidazole reductase that prevents the formation of nitroso radicals, the cornerstone of the antibacterial activity of metronidazole [29]. The nine nim genes that have been described to date are usually located on plasmids or chromosomes and can be transferred by conjugation [30]. Although important Gram-negative anaerobic periodontal pathogens such as Porphyromonas gingivalis have been shown to be susceptible to metronidazole [31], Aggregatibacter actinomycetemcomitans is inherently resistant due to its microaerophilic properties. Low proportions (<2%) of the subgingival microbiota from refractory periodontitis patients have been shown to be resistant to metronidazole [18, 32], while, even after adjunctive treatment with this antibiotic for 14 days, chronic periodontitis subjects had an increased proportion of, mainly, resistant Actinomyces spp., which returned to baseline levels after 90 days [25]. The findings of the current study which suggest that the nim gene was not detected in any of the investigated samples, regardless of the source (tongue, subgingival sample or deep pockets in periodontitis patients), are in agreement with the above mentioned reports and indicate that metronidazole is still an important antimicrobial compound for treating periodontal disease in Greek subjects. Furthermore, according to the personal national health records of the participants, metronidazole was not prescribed to participants in the course of the last 12 months, and a much lower percentage of subjects (18%) had been prescribed this antibiotic compared to β-lactams, during the last 5 years. The low frequency of metronidazole use could be another factor contributing to the absence of detecting the nim gene in any of the analysed samples.
Due to specific conditions in the oral cavity, there is a well-established biofilm organization of oral microbial communities, and the close proximity of its members allows for the exchange of genetic material between species and genera, as previously reported in the literature [33]. More than 1000 bacterial species have currently been identified in dental biofilm communities, twice as many as can be cultured [34]. Thus, only molecular techniques, as applied in the current study, could assess the potential for antibiotic resistance in these communities. The high frequency of detection of the blaTEM gene in the current study suggests the possibility of dissemination of resistance to other members in the polymicrobial community [35]. On the other hand, the methods used in the current study do not allow for identification of the bacterial sources of the blaTEM gene, and it should be noted that the presence of a given resistance gene does not guarantee that it will be expressed. For this purpose, function expression analyses such as proteomic or transcriptomic methodology should be applied. Future studies could apply, as mentioned, more sophisticated methodologies and in addition examine a variety of different genes conferring resistance to β-lactams such as SHV, AmpC and CTX already documented to encode for resistance in Enterobacteriaceae [36].
Taken collectively, data from the present study suggest the frequent presence, in the oral cavity, of the blaTEM gene encoding resistance to β-lactams, but not of the nim gene. Although the presence of a gene does not ensure its expression, it strongly suggests that antimicrobials should be judiciously used in treating periodontitis, especially in countries such as Greece, with high rates of antibiotic use and antimicrobial resistance.
CONCLUSION
The oral cavity of Greek subjects often harbours bacterial genes encoding for resistance to β-lactams. Thus, the antimicrobial activity of β-lactams, might be compromised. Since no metronidazole resistance genes were detected in the present study, its clinical use is supported by the current findings.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
This research has been partly co-financed by the European Union (European Social Fund - ESF) and Greek national funds through the Operational Program “Education and Lifelong Learning” of the National Strategic Reference Framework (NSRF) - Research Funding Program: Heracletus II. Investing in knowledge society through the European Social Fund.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publisher's Web site along with the published article.


