All published articles of this journal are available on ScienceDirect.
Association of Genetic Polymorphism of VAX1 (rs7078160 and rs4752028) with Non-Syndromic Oral Clefts: A Systematic Review and Meta-Analysis
Abstract
Background
Non-syndromic cleft lip with or without cleft palate (NSCL/P) is a prevalent craniofacial anomaly influenced by genetic and environmental factors. The VAX1 gene is crucial in craniofacial development, and its polymorphisms may contribute to the risk of NSCL/P.
Methods
A systematic review and meta-analysis of case–control studies was performed to evaluate the association of VAX1 polymorphisms (rs7078160 and rs4752028) with NSCL/P. Odds ratios (ORs) and 95% confidence intervals (CIs) were calculated under different genetic models. Subgroup, sensitivity, and publication bias analyses were also conducted.
Results
Seven studies comprising 1,483 cases and 1,519 controls were identified and included. For rs7078160, individuals with the AA genotype (homozygous for adenine) had a higher NSCL/P risk compared with GG genotype (homozygous for guanine) (OR = 2.34, 95% CI: 1.55–3.24), and AG carriers (heterozygous for adenine and guanine) were also at increased risk (OR = 1.31, 95% CI: 1.11–1.54). For rs4752028, CC (homozygous for cytosine) vs. TT (homozygous for thymine) (OR = 1.47, 95% CI: 1.09–1.97) and CT (heterozygous for cytosine and thymine) vs. TT (OR = 1.28, 95% CI: 1.01–1.52) were significantly associated with NSCL/P. Subgroup analyses showed stronger associations in non-Chinese populations, while results in Chinese cohorts were less consistent. Sensitivity analyses confirmed robustness for rs7078160, but funnel plot and Egger’s test indicated potential publication bias for rs4752028 (p=0.044).
Discussion
These results support the role of VAX1 polymorphisms in NSCL/P susceptibility, indicating possible ethnic differences and a polygenic inheritance pattern. However, limited sample sizes and publication bias for rs4752028 warrant cautious interpretation.
Conclusion
VAX1 rs7078160 and rs4752028 polymorphisms are significantly associated with increased NSCL/P risk, particularly among non-Chinese populations. Further large-scale multi-ethnic studies considering gene–environment interactions are warranted to validate these findings.
1. INTRODUCTION
Cleft lip with or without cleft palate (CL/P) is among the most prevalent congenital disorders in the craniofacial region, and with an incidence range from 1/1000 to 2/1000 live births depending on ethnic and racial populations [1, 2]. This congenital defect occurs in approximately 70% of cases without any other structural or developmental abnormalities and is classified as non-syndromic cleft lip with or without cleft palate (NSCL/P) [3]. Asians and Native Americans have the highest incidence of NSCL/P, whereas Africans have the lowest incidence. A cleft lip is the product of failed integration of the philtrum and maxillary prominences on one or both sides, while the failed merging of paired lateral palatine processes can lead to cleft palate [4-6]. Infants with NSCL/P may experience communication and feeding problems, psychiatric disorders, and middle ear infections. Although surgical repair and multidisciplinary care can improve function and appearance, these defects may still have long-term consequences for affected individuals, their families, and society [7]. This disease is a multifactorial disease, and environmental and genetic factors, such as alcohol and smoking, viral infections, nutrition, use of drugs and teratogens during pregnancy, are effective causes [8-10]. NSCL/P does not follow simple Mendelian inheritance, reflecting the complex interplay of multiple genes and environmental factors [11]. Genetic alterations can elevate the risk of this phenotype [12, 13]. Genome-wide association studies have recognized over 40 candidate genes and their related single-nucleotide polymorphisms (SNPs) [14-21]
The ventral anterior homeobox 1 (VAX1) gene, located on chromosome 10q25, is highly expressed during craniofacial development and plays a critical role in the formation of craniofacial structures. Several studies have reported associations between VAX1 variants and susceptibility to NSCL/P [22-27]. Based on the study by Mangold et al., conducted in European and Asian cohorts, the rs7078160 polymorphism of VAX1 was found to be associated with NSCL/P [28]. However, findings in the Chinese population have been inconsistent; for instance, Li et al. reported that the rs7078160 polymorphism of VAX1 was not associated with NSCL/P in Northern Han Chinese individuals [29].VAX1 plays an important role in craniofacial development and is considered a potential contributor to NSCL/P. Several studies have attempted to elucidate the relationship between the vax1 polymorphism and NSCL/P susceptibility in various racial and regional populations. These inconsistencies may be partly due to small sample sizes, which limit statistical power and reproducibility. To address the inconsistent results reported across different populations, we conducted a systematic review and meta-analysis to investigate the potential association between VAX1 polymorphisms and NSCL/P. This allows for the integration of available case–control studies, providing a more precise estimate of genetic risk. The main objectives of this work were to: (i) determine whether the commonly studied VAX1 variants (rs7078160 and rs4752028) are significantly associated with NSCL/P susceptibility, (ii) explore possible differences in genetic effects across diverse ethnic groups, (iii) evaluate the robustness and consistency of reported findings through sensitivity analyses, and (iv) assess potential publication bias and study quality. By clarifying the role of VAX1 in NSCL/P, our findings aim to improve understanding of its genetic basis, contribute to risk prediction, and support genetic counseling and preventive strategies.
2. METHODS
2.1. Search Strategy
This study protocol was prospectively registered in PROSPERO (CRD42024604186), and the systematic review and meta-analysis were conducted in accordance with the PRISMA guidelines [30]. We systematically searched PubMed, Scopus, Web of Science, Embase, and the Cochrane Library up to July 2025.These databases were selected because they are widely recognized, reputable, and collectively provide broad coverage of biomedical, genetic, and epidemiological research. Although some degree of overlap was anticipated, this combination minimized the risk of missing relevant studies while avoiding unnecessary duplication.
The research question was framed using the PICO (Population, Intervention / Exposure, Comparison, and Outcome) framework [31].
- The population consisted of individuals with non-syndromic cleft lip with or without cleft palate (NSCL/P).
- The exposure of interest was genetic polymorphisms of VAX1, specifically rs7078160 and rs4752028.
- The comparison group included unaffected or healthy controls without these polymorphisms.
- The outcome was the presence and strength of the association between VAX1 polymorphisms and susceptibility to NSCL/P.
The search strategy combined both MeSH terms and free-text words using Boolean operators. Reference lists of included studies and relevant reviews were manually screened for additional sources. To minimize publication bias, grey literature was also searched in OpenGrey, ProQuest Dissertations, Google Scholar, and relevant conference proceedings. Full details of the search strategy are presented in Appendix 1.
2.2. Eligibility Criteria
We included case-control studies that investigated at least one of the VAX1 polymorphisms (rs7078160 or rs4752028) in relation to NSCL/P, and that reported genotype frequencies or sufficient data to calculate odds ratios (ORs). Only human studies published in English were considered. Exclusion criteria were animal studies, case reports, reviews, editorials, commentaries, protocols, and studies assessing VAX1 polymorphisms in diseases other than NSCL/P. Articles lacking usable genotype data were also excluded.
2.3. Study Selection and Data Extraction
Two reviewers independently screened titles, abstracts, and full texts of potentially eligible studies. Any discrepancies were resolved through discussion or by consulting a third reviewer. Duplicate records were removed using EndNote with DOI/PMID matching and manual verification. The extracted data included the first author, publication year, country, sample size, investigated polymorphisms, and Hardy–Weinberg equilibrium (HWE) status in controls.
2.4. Risk of Bias Assessment
The methodological quality of the included studies was evaluated using the Newcastle–Ottawa Scale (NOS) [32]. This tool assesses three domains: the selection of study participants, the comparability of cases and controls, and the assessment of exposure or outcome. Studies were scored from 0 to 9 and classified as low risk of bias (7–9), moderate risk (4–6), or high risk (<4).
2.5. Statistical Analysis
Meta-analyses were performed using STATA version 14 [33]. Odds ratios with 95% confidence intervals were calculated under homozygous and heterozygous genetic models. A random-effects model (DerSimonian and Laird method) [34] was used as the primary approach due to anticipated clinical and methodological heterogeneity. At the same time, fixed-effects estimates were reported where heterogeneity was minimal (I2 < 50%). Between-study heterogeneity was quantified using the I2 statistic and tested using Cochran’s Q [35]. Subgroup analyses were conducted based on ethnicity (Chinese vs. non-Chinese), and sensitivity analyses were performed by excluding studies with small sample sizes (<300 participants) or those deviating from HWE. Publication bias was assessed using funnel plots and Egger’s regression test, with a p-value <0.05 considered evidence of asymmetry.
3. RESULTS
3.1. Study Selection and Characteristics
The initial search retrieved 475 records. After removal of duplicates and screening based on titles and abstracts, 14 full-text articles were assessed for eligibility. Following exclusions of letters, animal studies, and studies unrelated to VAX1 polymorphisms, seven studies [23, 29, 36-40] published between 2012 and 2025 were included in the quantitative synthesis (Fig. 1, PRISMA flow diagram). Among these, all seven studies evaluated the rs7078160 polymorphism, encompassing 1,483 cases and 1,519 controls, while five studies also examined rs4752028, including 1,073 cases and 1,151 controls. Four studies were conducted in China, and three in non-Chinese populations, including studies from Poland, Japan, and Saudi Arabia. Most studies reported significant associations between VAX1 polymorphisms and susceptibility to NSCL/P (Table 1).
| Author/Refs. | Year | Country | Design | Total Samples (Cases /Controls) | Investigated Polymorphisms | HWE P | Main Outcome |
|---|---|---|---|---|---|---|---|
| Sabbagh et al. [37] | 2019 | Saudi Arabian | Case-control | 360 (122/186) | rs7078160 rs4752028 |
0.13 0.154 |
There is a relationship between these two loci on 10q25 (rs4752028 and rs7078160) and NSCL/P in a population with high levels of consanguinity. |
| Li et al. [29] | 2017 | China | Case-control | 409 (186 /223) | rs7078160 rs4752028 |
0.7738 0.4263 |
VAX1 rs4752028 was weakly associated with NSCL/P development in the studied northern Chinese Han population |
| Niu et al. [39] | 2020 | China | Case-control | 784 (310/474) | rs7078160 rs4752028 |
0.125 0.06 |
SNPS of VAX1 rs7078160 increased NSCL/P risk in the Chinese population. |
| Mostowska et al. [38] | 2012 | Polish | Case-control | 412 (206 /206) | rs7078160 rs4752028 |
0.821 0.128 |
The association of polymorphisms located at chromosomal regions 10q25.3 and 17q22 with NSCL/P in the Polish population. |
| Peng et al. [23] | 2022 | China | Case-control | 311(249/62) | rs7078160 rs4752028 |
0.13 0.05 |
SNPS of VAX1 rs7078160 increased NSCL/P risk in the Chinese population. |
| Wang et al. [36] | 2020 | China | Case-control | 160(100/60) | rs7078160 | 0.43 | This study suggests that rs7078160 polymorphism is a risk factor of NSCL/P, and Vax1 is strongly associated with NSCL/P in Southern Chinese Han populations |
| Thao et al. [40] | 2025 | Japan | Case-control | 618 (310/308) | rs7078160 | 0.9987 | The VAX1 rs7078160 A allele was significantly associated with an increased risk for NSCL/P (OR = 1.67, p < 0.00001). |
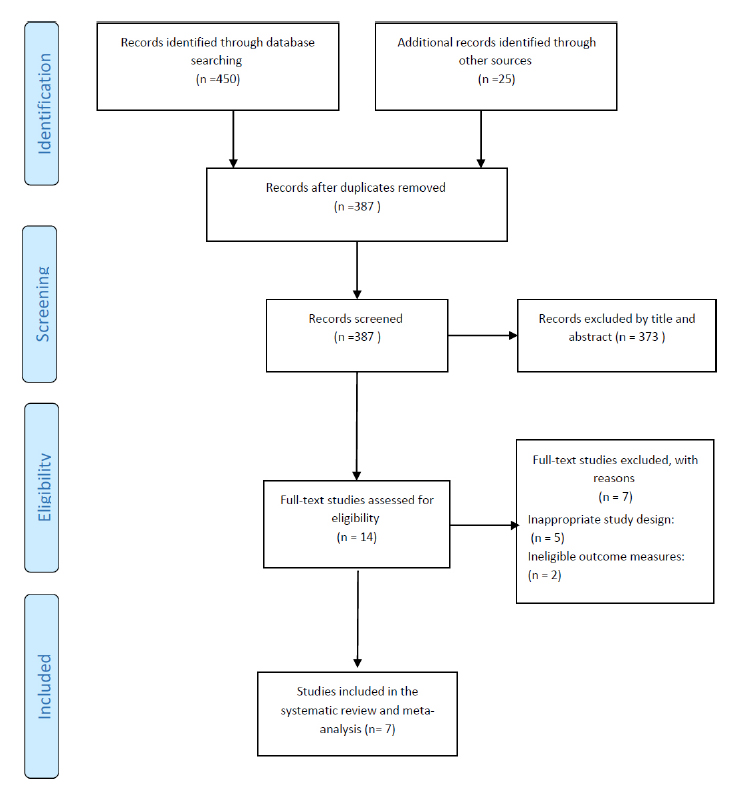
PRISMA flow diagram illustrating study selection process: from initial identification (n=475 records) to final inclusion (n=7 studies). PICO elements: Population (P): individuals with nonsyndromic cleft lip with or without palate (NSCL/P); Intervention/Exposure (I/E): presence of VAX1 rs7078160 or rs4752028 polymorphisms; Comparison (C): healthy controls; Outcome (O): association of VAX1 polymorphisms with NSCL/P risk
3.2. Risk of Bias
As shown in Table 2, based on the Newcastle–Ottawa Scale, four studies were classified as good quality (score ≥7) and three as fair quality (score 5–6). Importantly, none of the studies were judged as low quality (<5). Most investigations provided adequate case definitions, representative cases, and well-defined controls. Ascertainment of exposure was generally consistent across studies. However, some limitations were noted. Several studies did not fully adjust for potential confounders (e.g., sex distribution or maternal environmental risk factors), which may have influenced the observed associations. Despite these issues, the overall risk of bias was considered low to moderate, supporting the credibility of the pooled results.
| Author/Refs. | Selection | Comparability | Outcome | - | - | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Case Definition Adequate | Representativeness of the Cases | Selection of Controls | Definition of Controls | Main Factor* | Additional Factory | Ascertainment of Exposure | Same Method of Ascertainment for Cases and Controls | Nonresponse Rate | Total Score | Qualityz | |
| Sabbagh et al. [37] | + | + | + | + | + | - | + | + | - | 7 | Good |
| Li et al. [29] | + | + | + | + | + | + | + | + | - | 8 | Good |
| Niu et al. [39] | + | + | + | + | - | - | + | + | - | 6 | Fair |
| Mostowska et al. [38] | + | + | + | - | + | + | + | + | - | 7 | Good |
| Peng et al. [23] | + | + | + | - | + | - | + | + | - | 6 | Fair |
| Wang et al. [36] | + | + | + | - | + | + | + | - | - | 6 | Fair |
| Thao et al. [40] | + | + | + | + | + | + | + | + | - | 8 | Good |
y Sex was matched between 2 groups.
Z Good quality (score: >7) and fair quality (score: 5-7), Low quality (score: <5).
3.3. Meta-Analysis Findings
For rs7078160, individuals carrying the AA genotype (homozygous for adenine) had a significantly higher risk of NSCL/P compared with the GG genotype (homozygous for guanine) (OR = 2.34, 95% CI: 1.55–3.24, p <0.001). The heterozygous comparison (AG (heterozygous for adenine and guanine) vs. GG) also demonstrated a significant association (OR = 1.31, 95% CI: 1.11–1.54, p=0.001). Subgroup analyses revealed consistent associations in non-Chinese populations, where effect estimates were notably stronger than in Chinese cohorts. In Chinese populations, the homozygous model (AA vs. GG) remained significant, whereas the heterozygous model (AG vs. GG) showed no association (Fig. 2). Sensitivity analyses excluding studies with smaller sample sizes (<300 participants) did not alter the results, confirming the robustness of these findings (Fig. 3).

Forest plots of rs7078160 polymorphism under homozygous (AA vs. GG) and heterozygous (AG vs. GG) models, stratified by ethnicity.
Note: a). AA vs. GG in all studies (I2=57%, Random effect model).
b). AA vs. GG in non-Chinese population (I2=21%, Fixed effect model).
c). AA vs. GG in Chinese population (I2=3%, Fixed effect model).
d). AG vs. GG in all studies (I2=0%, Fixed effect model).
e). AG vs. GG in non-Chinese population (I2=0%, Fixed effect model).
f). AG vs. GG in Chinese population (I2=31%, Fixed effect model).

Sensitivity analysis for rs7078160 polymorphism showing stable results after exclusion of studies with small sample size (<300 participants).
Note: a). AA vs. GG (I2=63%, random effect model).
b). AG vs. GG (I2=65%, random effect model).
For rs4752028, the homozygous comparison (CC (homozygous for cytosine) vs. TT (homozygous for thymine)) showed a significant association with NSCL/P (OR = 1.47, 95% CI: 1.09–1.97, p=0.012). The heterozygous comparison (CT (heterozygous for cytosine and thymine) vs. TT) also indicated increased risk (OR = 1.28, 95% CI: 1.01–1.52, p=0.038). Subgroup analysis demonstrated that the associations were significant in non-Chinese populations, while results in Chinese cohorts were inconsistent, suggesting an ethnicity-specific genetic susceptibility. Considerable heterogeneity was observed for both comparisons of rs4752028, particularly in Chinese studies, which may explain the variability in effect estimates (Fig. 4).
Subgroup meta-analysis by NSCL/P subtype revealed that the rs7078160 (AA genotype) was significantly associated with an increased risk in the cleft lip only group, but showed no significant association in the cleft palate only subgroup, supporting its role as a potential susceptibility variant, mainly in cleft lip. For rs4752028, analysis was performed only in the cleft palate only subgroup, where no significant association was observed, indicating that its effect may be limited or context-dependent. These findings highlight the importance of considering NSCL/P subtypes when evaluating the contribution of VAX1 polymorphisms to craniofacial development (Fig. 5).

Forest plots of rs4752028 polymorphism under homozygous (CC vs. TT) and heterozygous (CT vs. TT) models, stratified by ethnicity.
Note: a). CC vs. TT in all studies (I2=77.78, Random effect model).
b). CC vs. TT in non-Chinese population (I2=0.00, Fixed effect model).
c). CC vs. TT in Chinese population (I2=58.13, Random effect model).
d). CT vs. TT in all studies (I2=60.46, Random effect model).
e). CT vs. TT in non-Chinese population (I2=76.46, Random effect model).
f). CT vs. TT in Chinese population (I2=20.1, Fixed effect model).
3.4. Publication Bias
For rs7078160, the funnel plot was symmetrical, and Egger’s test did not indicate significant publication bias (p = 0.134). For rs4752028, however, the funnel plot suggested asymmetry and Egger’s test confirmed the presence of publication bias (p = 0.044) (Fig. 6). These findings suggest that the observed effect for rs4752028 may be influenced by reporting bias and should be interpreted with caution, particularly given the limited number of available studies.
4. DISCUSSION
CL/P can lead to major aesthetic and functional problems, often resulting in significant dentofacial deformity. Patients with CL/P frequently present with Class III malocclusion, missing teeth, and maxillary deficiency [41], and approximately 25% to 35% require orthognathic surgery as a part of comprehensive treatment [42, 43]. This meta-analysis provides evidence linking VAX1 gene polymorphisms, specifically rs7078160 and rs4752028, to an increased risk of NSCL/P. The association was particularly significant in non-Chinese populations, underscoring the genetic heterogeneity observed across different ethnic groups. Notably, the low between-study heterogeneity (I2) observed for rs7078160 suggests a consistent effect across studies, enhancing the reliability of our findings.
4.1. Comparison with Existing Literature
Our results align with previous studies that identified VAX1 as a susceptibility gene for NSCL/P. For instance, Zhang et al. [44] reported a significant association between rs7078160 and NSCL/P in the Western Han Chinese population. Similarly, Li et al. [29] found an association between rs4752028 and NSCL/P in a Chinese cohort. However, studies in other populations have yielded inconsistent results. For example, a survey by Sabbagh et al. [37] found no significant association between VAX1 polymorphisms and NSCL/P in a Saudi Arabian cohort. These discrepancies may be attributed to differences in sample sizes, study designs, and genetic backgrounds.

Subgroup meta-analysis of VAX1 polymorphisms by NSCL/P subtype.
Note: a). AA vs. GG (rs7078160) in cleft lip only (I2= 0%, Fixed effect model).
b). AA vs. GG (rs7078160) in cleft palate only (I2= 0%, Fixed effect.
c). CC vs. TT (rs4752028) in cleft palate only (I2= 0%, Fixed effect model).

Funnel plots for (a) rs7078160 and (b) rs4752028. No evidence of publication bias was found for rs7078160, whereas slight asymmetry was observed for rs4752028 (Egger’s test, p=0.044), mainly due to the study by Niu Z et al., 2020 (China, 310 cases, 474 controls), which appeared as an outlier outside the funnel. This deviation is likely attributable to the study’s relatively large sample size and the magnitude of its effect estimate, which differed from the other included studies. Such characteristics can disproportionately influence the symmetry of the funnel plot, highlighting that the observed asymmetry may result from study-specific factors rather than true publication bias.
4.2. Genetic Heterogeneity and Subgroup Analyses
The observed ethnic differences in the strength of the association between VAX1 polymorphisms and NSCL/P risk highlight the importance of considering genetic heterogeneity in genetic association studies. Our subgroup analyses revealed a stronger association in non-Chinese populations, suggesting that genetic variants in VAX1 may have a more pronounced effect on NSCL/P risk in these groups. This finding is consistent with previous studies that have reported ethnic variations in the association between VAX1 polymorphisms and NSCL/P.
4.3. Insights from Genetic Inheritance Models
Recent work by Cheng et al. [6] emphasized that NSCL/P does not follow a simple Mendelian inheritance but rather fits within a spectrum ranging from monogenic to polygenic models. While rare variants in single genes such as IRF6, MSX1, and FOXE1 can contribute to clefting in specific families, most cases result from the cumulative effects of multiple common variants with modest effect sizes. Our findings regarding VAX1 add to this polygenic framework, supporting the notion that different loci jointly shape susceptibility. Moreover, Cheng et al. [6] highlighted the potential of using Polygenic Risk Scores (PRS) to integrate the effects of multiple SNPs, which could improve risk prediction and allow for more personalized approaches to genetic counseling and prevention strategies in NSCL/P.
4.4. Recent Genome-wide Association and Functional Studies
Recent large-scale genomic studies further support the role of genetic susceptibility in NSCL/P. For instance, Kumar et al. applied a massively parallel reporter assay (MPRA) in human oral epithelial cells. They demonstrated that several GWAS-identified SNPs, including those near IRF6, FOXE1, MAFB, TFAP2A, and TP63, exert functional effects on enhancer activity, thereby reinforcing their contribution to cleft pathogenesis [45]. Similarly, a recent multi-ethnic GWAS meta-analysis by Jia et al. identified 50 genome-wide significant loci, including 11 novel risk loci such as SHH, SOX9, PRICKLE1, and ALX1, highlighting both shared and ancestry-specific genetic mechanisms underlying NSCL/P [46]. Moreover, Curtis et al. showed that de novo variants located near GWAS-identified loci may play additional roles in orofacial cleft susceptibility, suggesting that integrating rare and common variants is essential for a comprehensive understanding of the genetic architecture of clefting [47].
5. LIMITATIONS
Several limitations should be considered when interpreting our findings. First, the inclusion of only published studies in English may have introduced publication bias. Future meta-analyses should aim to include studies published in languages other than English to provide a more comprehensive assessment of the association between VAX1 polymorphisms and NSCL/P risk. Second, most of the included studies did not account for maternal environmental factors, such as smoking and alcohol consumption during pregnancy, which could confound the observed associations. Cheng et al. [6] also stressed that gene–environment interactions are critical, as environmental exposures may modulate the penetrance of genetic variants. Third, some studies had relatively small sample sizes and limited ethnic diversity, which may reduce statistical power and generalizability. Finally, most studies did not stratify cases by NSCL/P subtype, which limits insight into the subtype-specific effects of VAX1 polymorphisms.
5.1. Clinical and Public Health Implications
Understanding the genetic factors contributing to NSCL/P can inform risk assessment and preventive strategies. Identifying individuals with VAX1 polymorphisms associated with an increased risk of NSCL/P could facilitate early interventions and targeted genetic counseling. Furthermore, polygenic models such as PRS, as highlighted by Cheng et al. [6], may provide a valuable tool for stratifying patients by genetic risk and developing tailored preventive or therapeutic approaches.
6. FUTURE RESEARCH DIRECTIONS
Future studies should aim to replicate our findings in diverse populations to confirm the association between VAX1 polymorphisms and the risk of NSCL/P. Additionally, functional studies are needed to elucidate the biological mechanisms by which VAX1 variants influence craniofacial development. Investigating gene-environment interactions, particularly maternal exposures during pregnancy, will provide a more comprehensive understanding of the etiology of NSCL/P.
CONCLUSION
These findings demonstrate a significant association between VAX1 polymorphisms (rs7078160 and rs4752028) and the risk of NSCL/P. They underscore the role of genetic factors in the etiology of NSCL/P and suggest that identifying high-risk individuals could support early intervention and personalized management. Further research is needed to clarify the mechanisms by which VAX1 variants influence craniofacial development and to explore potential preventive or therapeutic strategies. In line with Cheng et al. [6], our results strengthen the evidence that NSCL/P arises from complex polygenic inheritance patterns, and future application of PRS may enhance clinical translation in risk prediction and genetic counseling.
AUTHORS’ CONTRIBUTIONS
The authors confirm contribution to the paper as follows: P.M. and Z.M.: Contributed to the conception and design of the study; P.M., Z.M., and F.S.: Involved in screening and evaluating the literature; F.S. and Z.M.: Analyzed the data and drafted the manuscript; P.M.: Revised and critically reviewed the manuscript. All authors approved the final version of the submission.
LIST OF ABBREVIATIONS
| CL/P | = Cleft lip with or without cleft palate |
| NSCL/P | = Non-syndromic cleft lip with or without cleft palate |
| VAX1 | = Ventral Anterior Homeobox 1 |
AVAILABILITY OF DATA AND MATERIALS
The data supporting the findings of the article will be available from the corresponding author [F.S] upon reasonable request.
ACKNOWLEDGEMENTS
Declared none.
SUPPLEMENTARY MATERIAL
PRISMA checklist is available as supplementary material on the publisher’s website along with the published article.


